Professors from Chemical Engineering and Computer Science Use Big Data to Design New Nanomaterials
Thanks to a $1.4 million three-year grant from the U.S. Department of Energy, Sanat Kumar, Professor and Department Chair, Chemical Engineering, Venkat Venkatasubramanian, Samuel Ruben-Peter G. Viele Professor of Engineering, and Michael Collins, Vikram S. Pandit Professor of Computer Science, are applying big data concepts and techniques to discover and design new nanomaterials—a priority area under the White House's Materials Genome Initiative—using a methodology that could revolutionize materials design.
The Columbia team is collaborating with Oleg Gang of Brookhaven National Laboratory on “DNA-Grafted Building Blocks Designed to Self-Assemble into Desired Nanostructures,” on what they say is the first collaborative research project between the Engineering School’s chemical engineering and computer science departments, and, adds Kumar, “one of the first data-intensive grants that came to Columbia Engineering after the formation of the Institute for Data Sciences and Engineering.”
"There is an increasing demand, especially in the chemical and pharmaceutical industries, for the design of new materials and formulations with specialized physical, chemical, and biological properties,” says Venkatasubramanian. “This search for novel materials has become an essential part of these industries’ R&D and presents a highly complex but critical challenge for us to solve, as these materials encompass a wide variety of products impacting daily life, from drugs and agricultural chemicals such as pesticides or herbicides, to refrigerants, fuel additives, paints and varnishes, and even personal care products such as shampoo.” He notes that while the rapidly growing avalanche of high throughput experimentation data has created exciting opportunities, the data also present a major modeling and big data informatics challenge for materials design and discovery. In addition, the traditional trial-and-error approaches currently used for materials design are extremely time-consuming and expensive, and can not only cause delays in the time-to-market period but also miss potential solutions.
“Guided by pioneering work by Ned Seeman at NYU and Chad Mirkin at Northwestern, we now recognize that the use of DNA provides us with a natural tool for directed particle assembly,” Kumar says. “DNA assembly into double helices is chemically specific—particles with short single-stranded DNA grafted on their surfaces will be bridged together only if those strands have complementary base sequences.” To date, researchers have used DNA-grafted particles to form four different crystal structures, and the crystal data bank has several hundred known crystal structures. The open question, as Kumar observes, is whether they can a-priori design DNA-grafted particles that spontaneously assemble into not only these four structures, but into any desired structure in the databank.
The team plans to develop computer-based tools to design DNA-grafted colloidal building blocks (DBB) that will spontaneously and reliably assemble into desired structures. These structures are not only useful for making materials with controllable photonic band gaps, but also for developing metamaterials with variable indices of refraction. Once created, these materials could then be integrated into chips and advance the future of computers and telecommunications. Instead of doing this through the conventional Edisonian or “forward” model of research where building blocks—in this case the DNA-grafted particles—are first synthesized and then examined as to the structures they might assemble into, the team proposes to create a research model in which a building block is designed in such a way that it will spontaneously assemble into a desired structure.
“Very often, scientists will make a new material and then think ‘What can I do with this?’,” Kumar explains. “We want to reverse this way of thinking—we want to first catalogue the things we want the material to do or be, and then figure out a way to make it. The analogy here is with a commodity item like a car or a TV. For example, with a car we know consumers want 4-wheel drive, ABS, a GPS system, and so on. They tell us what they want and then we engineers figure out how to design and put the pieces together. With this new research, we hope to develop methods that will enable us to pick the formulations we desire and design nanoparticles that can assemble into the structure we want. This could radically transform the field of product design.”
The research team has strong complementary experimental and theoretical skills. Kumar has developed and implemented highly efficient methods that allow you to take the molecules you want, and using concepts no more complicated than the Newton’s laws, predict what structures they will assemble into. Venkatasubramanian is an expert in molecular design strategies and the use of mathematical optimization and big data analytical techniques that are essential for successfully implementing the underpinning product design concepts. Collins is a specialist in large-scale data-mining and machine learning algorithms that go beyond conventional classification problems in their prediction of complex structures such as sequences, and Gang is a pioneer in experimentally assembling DNA-grafted colloids into ordered structures and in characterizing these assemblies using a suite of unique experimental tools available at the Center for Functional Nanostructures at Brookhaven.
“It is this diverse set of tools that enables us to begin to put together the design platform, one that we believe will span not only this particular problem, but more broadly the field of chemical product design,” Kumar observes.
“Drs. Kumar and Gang and I have recently begun to collaborate on understanding the factors that drive DNA-grafted nanoparticles to assemble into ordered arrays,” says Venkatasubramanian. “Adding the critical design component we propose to do with this DOE grant will enable our team to start focusing our studies on systems of particular interest in the context of applications.”
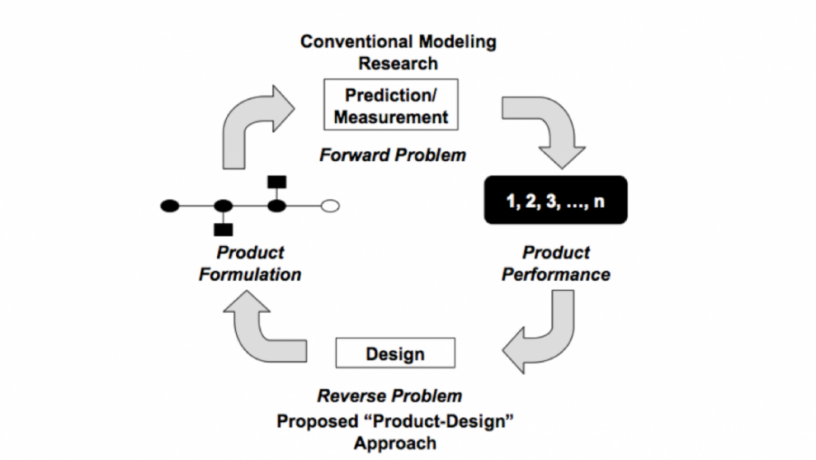
Comparison of conventional and proposed paradigms.